2019-04-02
Personalized nutrition is emerging as a field of interest due to an increase in coverage in the general media and the growing awareness that no two people can have identical responses to the same meal. One research paper that detailed the unique postprandial glycemic response (PPGR) to dietary intake was published by Dr. Segal and Dr. Elinav at the Weizmann Institute in 2015. In the past, this variable response from person to person has been identified however the specific factors behind this variability are described for the first time in this paper. Currently, to achieve a homeostatic glycemic balance, carbohydrate intake is primarily taken in to consideration. The Segal lab took this a step further and created an algorithm based on blood parameters, anthropometrics, physical activity, lifestyle behaviours and the gut microbiome which allows for individual predictions of PPGRs in response to different foods.
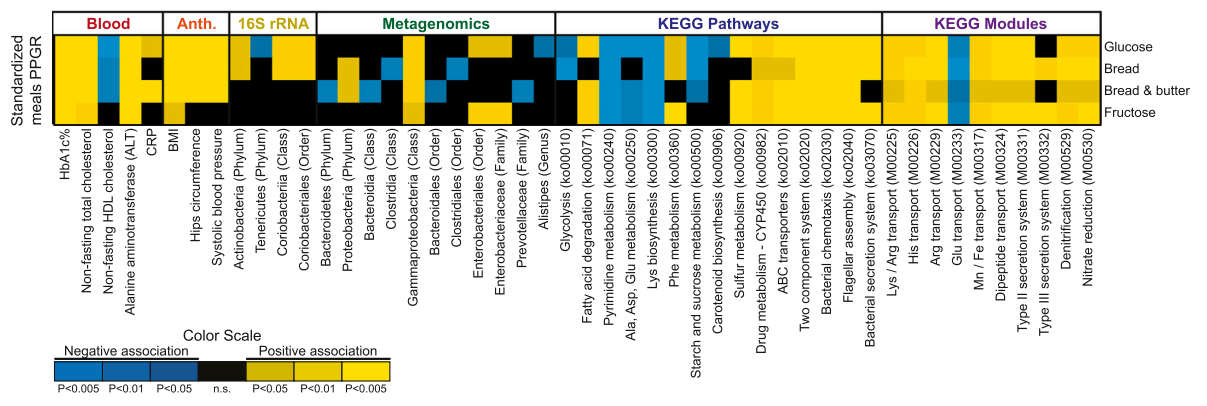
Fig 1. High Interpersonal Variability in the Postprandial Glycemic Response to the Same Meal. Heatmap (subset) of statistically significant associations (p < 0.05, FDR corrected) between participant’s standardized meals PPGRs and participants’ clinical and microbiome data.
Figure 2 below outlines the associations between gut microbiota and standardized meal PPGRs among other clinical factors. It is interesting to see how certain types of foods elicit a higher glycemic response in the presence of certain bacteria. Knowing everyone has a unique gut microbial blueprint, this highlights the truly diverse response we all have to particular foods. This is useful in the prediction of predisposition to type II diabetes mellitus (TIID) in patients. Patients with certain populations of bacteria may respond with higher or lower PPGRs than others, thus making it difficult to assert patient risk to TIID without proper microbiome assessments.
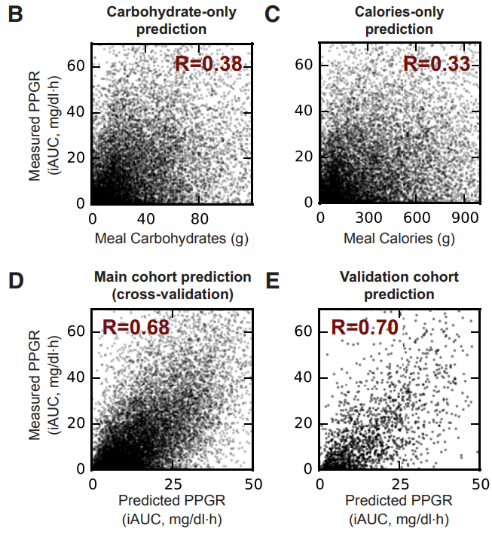
Fig 2. Accurate Predictions of Personalized Postprandial Glycemic Responses. (B-E) PPGR predictions. Dots represent predicted (x axis) and CGM-measured PPGR (y axis) for meals, for a model based: only on the meal’s carbohydrate content (B); only on the meal’s Caloric content (C); our predictor evaluated in leave-one-person-out cross validation on the main 800-person cohort (D); our predictor evaluated on the independent 100-person validation cohort (E); Pearson correlation of predicted and measured PPGRs is indicated.
Since standard PPGRs are predicted using caloric and carbohydrate measures, these two parameters were tested against the algorithm developed to confirm validity. As seen in figure 2B, C, D, and E, through an R validation measure, the authors were able to confirm that the algorithm introduced an accurate way of predicting PPGRs with the validation cohort resulting in an R value of 0.70 compared to the carbohydrate at 0.38, and calorie at 0.33. Based on these predictions it is reiterated how factors, such as gut microbiota, are essential in the prediction of PPGRs and how calorie and carbohydrate based predictions are insufficient. In an era where hyperglycemia is omnipresent, the importance of getting the full picture of our response to glycaemia rich diets is highlighted. The authors used DNA Genotek’s OMNIgene·GUT stool collection and stabilization kits for this predictive study to prevent the progression of TIID in patients predisposed to it. Having created a collection kit that stabilizes the microbial profile from stool samples at ambient temperatures for up to 60 days, DNA Genotek has made it much easier to collect samples from donors at home and in remote geographical regions.
Finally, a heat map was established in figure 6C based on 26 participants that were subject to two, one-week diets: one “good” diet consisting of foods lowering PPGRs, and one “bad” diet increasing PPGRs. These diets were determined through the use of the algorithm to tailor the predicted “good” and “bad” based on different parameters including the gut microbiome. Every participant has a personalized expression of certain bacteria. The bacteria associated with good and bad diets are consistent and many show significant increased or decreased presence in response to diet. Certain bacteria that are present in lower levels in TIID are seen to increase with the “good” diet, as well as high levels of other bacteria associated with TIID were seen to decrease. This further accentuates the truly unique PPGRs from person-to-person depending on the presence of certain microbiomes, once again confirming the importance of gut microbiome studies in the prediction of TIID predisposition. Additionally, good diets seem to cause long-term change in microbial populations, which provides an additional benefit of having tailored diets.
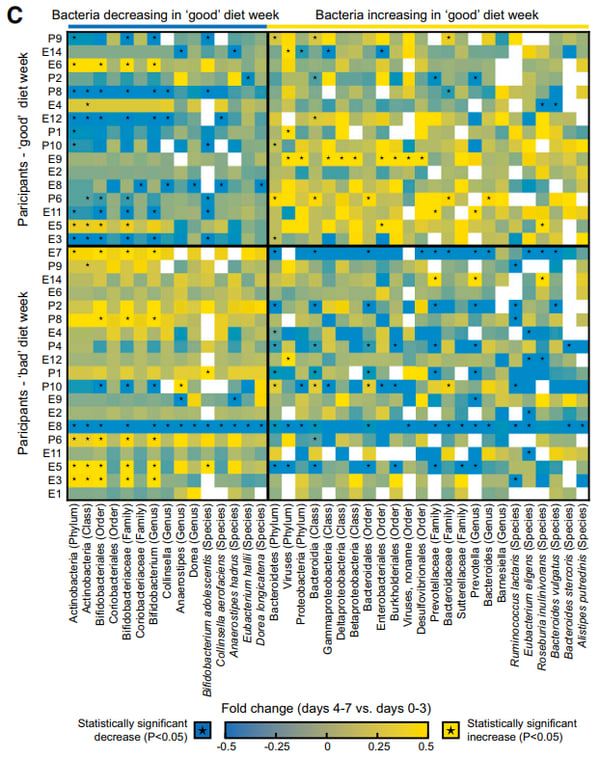
Fig 3. Dietary Interventions Induce Consistent Alterations to the Gut Microbiota Composition (C) Heatmap of taxa with opposite trends of change in RA between ‘‘good’’ and ‘‘bad’’ intervention weeks that was consistent across participant and statistically significant (Mann-Whitney U-test between changes in the ‘‘good’’ and ‘‘bad’’ weeks, p < 0.05, FDR corrected). Left and right column blocks shows bacteria increasing and decreasing in their RA following the ‘‘good’’ diet, respectively, and conversely for the ‘‘bad’’ diet. Colored entries represent the (log) fold change between the RA of a taxon (x axis) between days 4–7 and 0–3 within each participant (y axis). Asterisks indicate a statistically significant fold change.
What contributes to our microbiome?
During a webinar hosted by The Scientist, Dr. Eran Segal enlightened us on the different contributing factors affecting our microbiome. Studies conducted by his lab concluded that ancestry and SNPs had no significant association with the gut microbiome. Approximately 20% of our microbiome beta-diversity is associated with environmental factors such as diet, drugs, activity, smoking, and quality of life. It is interesting to hear that although genetics are not associated with microbial content, microbiome and genetics have an additive effect on the predicted phenotype. Genetics, microbial profile, age, gender, and calories can variably contribute to phenotypes. Personalized nutrition can be key to determining what happens inside our bodies, and this research indicates that it is not simply genetics, but also the microbes that live in our gut that make us who we are.
For more information on our OMNIgene·GUT microbial collection devices, please do not hesitate to contact us at info@dnagenotek.com.