2019-02-12
One of the many ways to ensure optimal isolation of bacterial DNA from stool samples, is through the use of an efficient extraction methodology. Our customers have been asking for extraction recommendations, so we conducted a validation study comparing four different manual extraction methods. There are many steps during the DNA isolation process where a multitude of biases can be introduced. The OMNIgene•GUT device reduces biases during stool sample collection to provide our users with an accurate snapshot of the in vivo gut microbial DNA content at the time of collection. This snapshot is maintained at ambient temperature for 60 days making it flexible and reliable to use when you are ready to extract microbial DNA.
That being said, the next step is to carry this accurate snapshot forward through to extraction and analysis. We want to help you choose the most efficient extraction method, thus we carried out a direct comparison between the QIAamp PowerFecal DNA kit, QIAamp PowerFecal Pro DNA kit, RBB+C (repeated bead beating and column), and ZymoBIOMICS DNA Miniprep kit. A summary of the four extraction methods and their respective performance on yields, lysis, DNA purity and integrity can be found in Table 1 below:
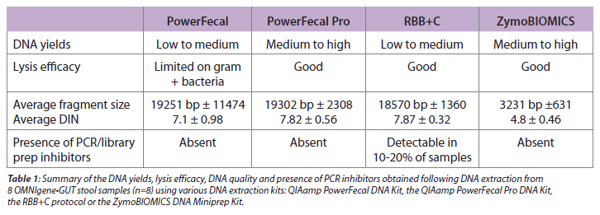
Based on our findings, the best performing microbial DNA extraction method for OMNIgene•GUT is the PowerFecal Pro extraction method. Our results indicate that the PowerFecal Pro efficiently lysed cells (including tough to lyse gram-positive bacteria) while maintaining DNA integrity/purity and high yields. This extraction method is also effective for low biomass samples such as infant stool. It can be used on all samples types of the Bristol scale, which makes it more versatile for large population studies.
The other three extractions methods showed limitations in at least one of the performance parameters tested making them sub-optimal compared to PowerFecal Pro.
What are the implications for your microbiome studies?
Microbiome studies depend highly on the accurate measurement of microbial community diversity present in a sample at a given time. Anything that will hinder this leads to unreliable results and creates the need for repetition of experimental procedures, which is generally not feasible.
Using an extraction method that cannot efficiently lyse gram-positive bacteria, will bias results and skew data toward gram-negative bacteria, and ultimately prevent the researcher from obtaining a true representation of the microbial profile.
Additionally, recovery of DNA of low purity or integrity is likely to make it difficult to perform certain next generation sequencing techniques such as long range sequencing or require additional sample clean up prior to analysis. This results in obstacles when trying to prepare libraries for various sequencing applications, and thus requiring researchers to either re-sample and/or re-extract for better results.
What does this mean for your research?
Considering all of the implications in using a sub-par extraction methodology, it is worth carefully selecting the best collection and extraction kit combination. This will ensure that you will not need to invest more time and money into re-extracting and/or potentially re-collecting stool samples to obtain desirable results. Using a sub-par experimental design will undoubtedly slow down the overall study progress, increase costs, and deplete resources. Thus, for optimized efficiency when it comes to DNA quality, best results are achieved using a combination of our OMNIgene•GUT kit and the PowerFecal Pro extraction method.
For more information on the experimental findings of this comparison, you can view the application note. If you would like to evaluate our OMNIgene•GUT stool collection kits, please email us at info@dnagenotek.com.