2019-05-01
Most people consider the nature of collecting fecal samples unpleasant and inconvenient. One of the challenges for the researcher conducting a gut microbiome study, is to provide the participants with a comprehensive and simple collection procedure without compromising the integrity of the sample. There is a mountain of data on collection methods, temperature holds, stabilizers vs fresh, individual chemistries, effects of freezing on viability, and DNA integrity…how can we make sense out of this often conflicting data?
In this second segment to our three part cold chain stool sample collection series (click to read part 1), we explore where error and bias can be introduced when using cold-chain sample collection and transportation of stool samples. We look at two important factors regarding stool collection for microbiome research: simplicity and stability.
Simplicity
Simplicity is critical to achieve high donor compliance and participation. In addition, ensuring that samples are collected properly is the first step to achieving bacterial stability and reproducibility between samples.
Donor compliance – collection instructions and user protocols
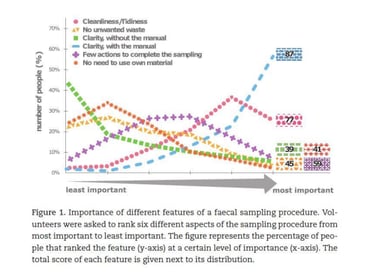 To prevent error in stool sampling and bias in the stool derived bacterial profile, it is important to have a simple and clear collection protocol. The Flemish Gut Project, from Vlaams Instituut voor Biotechnologie (VIB), found that the most important aspect to donor compliance is clear and simple instructions (Figure 1). They evaluated six different aspects of a sample collection and concluded that donors valued clear and simple instructions overall when it came to the collection procedure. [1]
To prevent error in stool sampling and bias in the stool derived bacterial profile, it is important to have a simple and clear collection protocol. The Flemish Gut Project, from Vlaams Instituut voor Biotechnologie (VIB), found that the most important aspect to donor compliance is clear and simple instructions (Figure 1). They evaluated six different aspects of a sample collection and concluded that donors valued clear and simple instructions overall when it came to the collection procedure. [1]
Donor compliance – A case study using FTA cards
In addition to cold-chain sample collection, some researchers use FTA cards (Flinders Technology Associates cards), which are paper made products laced with a mixture of chemicals that lyse cells and stabilize nucleic acids when the stool sample is smeared onto the card.[2] The Knight Lab from the University of California San Diego (UCSD), tested the effects of FTA cards in a lab setting with trained personnel and infield with inexperienced donors.
- Lab simulated: they reported that both FTA cards and OMNIgene•GUT resulted in similar fecal communities with compositional changes similar to those observed between technical replicates. However, this was performed in a lab setting with trained personnel and a controlled environment. [3]
- Field collected: they reported that the FTA cards preserved samples with the least similar microbial composition and abundance compared to fresh samples when done infield with inexperienced donors and an uncontrolled environment (see Figure 2). [4]
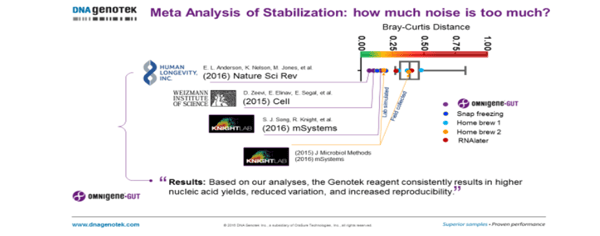 Figure 2. Comparing the effect of stabilization and inexperienced vs scientifically trained donors on the microbial profile (DNA Genotek). The two papers from the Knight Lab are highlighted in this figure. The lab simulated Bray-Curtis distance is close to those found with OMNIgene·GUT whilst the field collected samples cluster with those from RNA later and home brew methods which have questionable efficacy.[2],[3]
Figure 2. Comparing the effect of stabilization and inexperienced vs scientifically trained donors on the microbial profile (DNA Genotek). The two papers from the Knight Lab are highlighted in this figure. The lab simulated Bray-Curtis distance is close to those found with OMNIgene·GUT whilst the field collected samples cluster with those from RNA later and home brew methods which have questionable efficacy.[2],[3]
Complex collection methods such as cold-chain and FTA cards may work in the lab with trained personnel but once the study begins and the collection is done with untrained donors, the samples collected may not produce accurate and reproducible results.
The importance of stool sample stability
The most important part of stool sample collection is that the microbial profile remains stable from the moment of collection through to lab processing. Traditionally, stool samples would be frozen for transport to the lab, slowing bacterial growth and limiting deviations from the original bacterial composition.
From collection to the lab, there are 3 main periods of temperature fluctuation:
- Time of collection to placement in freezer (room temperature)
- Time in household freezer (-20ºC to -2ºC, cycling every 24 hours)
- Time spent in shipping (variable temperature range)
None of these temperature conditions are ideal for bacterial stability; therefore, using this method will likely result in profile changes and biased samples.
Stability in a participant’s home
It is difficult to characterize change in the bacterial profile that may occur from the time a sample is collected until it is placed in the freezer. Gorzelak et al 2015, suggested that significant profile changes can occur during this time frame and thus recommends that participants freeze their samples within 15 minutes. Unfortunately, household freezers are not built like lab-grade freezers.[5]
“In studies where human participants are asked to self-sample and ship to a laboratory, for instance, samples may experience temperature fluctuations during storage (as home freezers undergo automatic defrost cycles). In regions with extreme temperatures or in cases of freezer failure, samples may additionally be exposed to extreme heat.” [3]
Therefore, placing samples in home freezers with temperature fluctuations may affect the accuracy of downstream DNA based bacterial taxa detection.[5] With lab grade freezers there is data to show that even when unstabilized samples are stored at -20°C, the bacterial profile is not stable; therefore, it is recommended to store unstabilized samples at -80ºC rather than at -20ºC for long-term storage (see Figure 3 below).
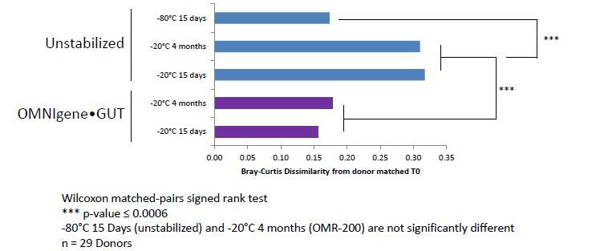 Figure 3. Unstabilized samples held at -20ºC (lab grade freezer) for 15 days are significantly dissimilar to donor paired samples extracted at t=0 (DNA Genotek).
Figure 3. Unstabilized samples held at -20ºC (lab grade freezer) for 15 days are significantly dissimilar to donor paired samples extracted at t=0 (DNA Genotek).
Shipping stability
The period of time a stool sample spends in transit can range from 1-7 days. [6]
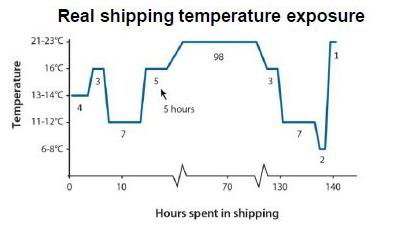 Figure 4. Hours spent in shipping at corresponding temperatures (DNA Genotek).
Figure 4. Hours spent in shipping at corresponding temperatures (DNA Genotek).
At DNA Genotek, we ran a study to test the temperature control efficiency of the collection container (Figure 4). This protocol involved placing fecal samples in a Styrofoam container with frozen gel packs and ensuring that processing occurred within 24 hours. 23 donors collected fecal samples and mailed their samples to a lab. The temperature inside the box was measured upon arrival and was <5°C in 40% of the samples after 1 day in the mail, 65% after 2 days and 100% after 3 or more days in the mail.
NOTE: 1 box arrived at ambient temperature after only 1 day in transit, which indicates that the donor did not comply with the sample collection instructions and did not completely freeze the icepacks prior to shipping the sample.[7] Therefore, due to the fluctuating temperatures that fecal samples are exposed to during cold-chain transportation, this method is not effectively preventing bacterial growth, potentially introducing a bias in the microbial profiling.
Conclusion
We discussed compliance issues for gut microbiome studies and also discussed the difficulty in recruiting and keeping participants engaged. We highlighted the importance of collection simplicity and sample stability. In addition, we provided data to demonstrate that using OMNIgene·GUT is an easy, and reliable stool collection method for gut microbiome studies.[8]
In our final blog of our 3 part series, we’ll look at additional biases that can be introduced into a work flow from extraction through to sequencing and how we believe a true validation should be performed to get the best results from your study.
For more information on our OMNIgene·GUT microbial collection devices, please do not hesitate to contact us at info@dnagenotek.com.
Links to other blogs in this series
Cold chain stool sample collection - as series of unfortunate events part 1 of 3
Cold chain stool sample collection - as series of unfortunate events part 3 of 3
References:
[1] Vandeputte et al., 2017. Practical considerations for large scale gut microbiome studies. FEMS Microbiology Reviews. 41(1): S154-S167 (2017).
[2] https://bitesizebio.com/9350/the-pros-and-cons-of-storing-dna-on-cards/
[3] Song et al. Preservation Methods Differ in Fecal Microbiome Stability, Affecting Suitability for Field Studies
mSystems 1(3)., DOI: 10.1128/mSystems.00021-16 (2016).
[4] Amato, K. An introduction to microbiome analysis for human biology applications. Am J Hum Biol. 29 (1). DOI: 10.1002/ajhb.22931 (2017).
[5] Gorzelak et al, 2015. Methods for Improving Human Gut Microbiome Data by Reducing Variability through Sample Processing and Storage of Stool. PLoS ONE 10(8): e0134802.
[6] DNA Genotek poster: https://www.dnagenotek.com/ROW/pdf/Identification-of-factors-impacting-reproducibility-and-quality-of-microbiome-profile-analysis-MK-00464.pdf
[7] DNA Genotek poster: https://www.dnagenotek.com/ROW/pdf/Critical-To-Quality-pre-analytical-factors-and-their-impact-on-microbiome-analysis-MK-00629.pdf
[8] Anderson, E.L. et al. A robust ambient temperature collection and stabilization strategy. Enabling worldwide functional studies of the human microbiome. Sci. Rep. 6, 31731; doi: 10.1038/srep31731 (2016).